The capsid of Adeno-associated Virus (AAV) is a naturally occurring, replication-deficient, virus that is widely considered the frontrunner for solving the delivery problem in gene therapy. These viruses are known to be harmless to humans, and are relatively simple to manipulate. One well-known drawback of natural capsids however, which are currently used for delivery, is that many patients with pre-existing immunity to the virus (due to previous natural exposure) may be ineligible for life-changing treatment.
In previous work (published in Science), we validated the use of computational models in conjunction with high-throughput experiments to design better liver-targeting variants of naturally occurring AAV capsids. In that work we were primarily focused on single edits to the capsid, and hypothesis that the effect of a combination of single mutations, at least when the number of total edits is limited, can be approximated by the sum of the effect of each mutation. Through this approach we validated that a model-guided method can lead to more efficient design of better capsids for more effective liver targeting.
The paradigm of measuring the effects of mutations independently and combining the best ones no longer works as we attempt to modify the capsid beyond a handful of mutations. Making capsids with many changes relative to natural variants increases our chances of being able to treat the thousands of potential recipients of gene therapy by evading pre-existing immunity. To achieve the ability to introduce a large number of changes to the capsid sequence without breaking its essential abilities, a wholly new approach was needed, which our latest study in Nature Biotechnology aims to address. Our goal was to design highly diverse AAV capsids, for which we used much more advanced machine learning models and trained on more complex datasets The work was a result of years of collaboration between teams at Dyno, Harvard’s Wyss Institute, and Google Research.
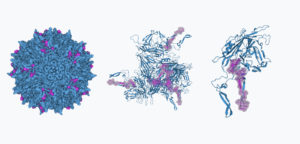
To test these methods, we focused on a representative region of the capsid (positions 560-588, seen in pink in the fully assembled virus, the hexamer assembly, and the individual subunit in the figure above) that had both surface-exposed and buried residues (Generally speaking, surface-exposed residues are known to be more mutation-tolerant) This region is also well known for the presence of immunogenic structures, as well as its role for tissue targeting. Our aim was to introduce as many mutations as we could in this 28 amino-region, including substitutions and insertions, the latter of which is a less common type of mutation in nature. When we started this study, it was unknown if machine learning models would be reliable for predicting the effects of mutations for variants beyond 5-10 edits to the original sequence. We expected this was possible, however, based on analyzing the diversity of sequences that have been isolated from natural sources. In this region, the average difference between two AAV serotypes is 12 amino-acids (often with few or no insertions). Nonetheless, we pushed the models to propose sequences with up to 29 substitutions and insertions.
Using the naturally observed level of diversity as a benchmark, we set our goal to generate diversity beyond that observed in nature, while maintaining the capsid’s viability. After screening billions of potential sequences in-silico using machine learning models, we settled on ~200,000 designed variants which we experimentally tested for their viability. Of those, approximately 110,000 produced viable viruses (many of our attempts were deep into the sequence space, where it is very hard to propose viable viruses). About 57,000 variants were farther than 12 mutations away from the AAV2 serotype. By generating more than two thousand sequences that were 25 or more mutations away, we decisively demonstrated the power of machine learning models to design diverse synthetic capsid sequences.
In this study, largely conducted before Dyno’s official launch, we report one of the largest AI-driven protein design assays published to date and validated the utility of these techniques for capsid engineering. The success of this approach bolstered our confidence in Dyno’s foundational science. Building upon this foundation, we have established infrastructure and machine learning techniques at Dyno to expand and optimize the AAV repertoire for multiple traits (including in-vivo targeting of challenging tissues), multiple serotypes, and at a larger scale. This study is just the beginning of our endeavour.
This work was a multi-year collaboration between Dyno co-founders Eric Kelsic, Sam Sinai, and George Church, colleagues at Harvard’s Wyss Institute including Nina Jain and Pierce Ogden and members of the Google Accelerated Science team including Patrick Riley, co-first authors Drew Bryant, Ali Bashir, and co-corresponding author Lucy Colwell.